Enzyme Kinetics: Velocity - Purdue University
ENZYME KINETICS: • The rate of the reaction catalyzed by enzyme E A + B ↔ P is defined as -Δ[A] or -Δ[B] or Δ[P] Δt Δt Δt • A and B changes are negative because the substrates are disappearing • P change is positive because product is being formed. • Enzyme activity can be assayed in many ways
Download Enzyme Kinetics: Velocity - Purdue University
Information
Domain:
Source:
Link to this page:
Please notify us if you found a problem with this document:
Documents from same domain
20.1 Mass Spectrometry Fundamentals - Purdue …
www.chem.purdue.edu20.1 Mass Spectrometry Fundamentals ... The mass spectrum can be used to determine molecular structure, to quantify single compounds or mixtures of
Fundamentals, Mass, Molecular, Spectrometry, 1 mass spectrometry fundamentals
Ch. 7 - Enolates - Purdue University
www.chem.purdue.edu651.06 209 Enolates Enolates are the conjugate anions of carbonyl compounds. Although they have been known and used since the turn of the 20th Century, it was the development of “specific
PURDUE UNIVERSITY INSTRUMENT VAN PROJECT
www.chem.purdue.eduFACTORS AFFECTING ENZYME ACTIVITY PURDUE UNIVERSITY INSTRUMENT VAN PROJECT QUESTIONS 1. In what way is the rate of the reaction dependent on the concentration of the enzyme?
Project, University, Activity, Rates, Instruments, Reactions, Purdue, Purdue university instrument van project, Activity purdue university instrument van project
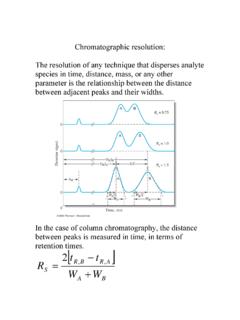
how to determine HETP from a peak - Purdue University
www.chem.purdue.eduChromatographic resolution is defined as the ratio of the difference of the retention times of two components to the sum of the peaks widths at the
The Fermentation of Pyruvate - Purdue University
www.chem.purdue.eduundergo lactate fermentation. Two pyruvates are converted to two lactic acid molecules, which ionize to form lactate. In this process two NADH + H+ are converted to two NAD+. Our muscle cells can undergo this process when they are in oxygen debt. If enough oxygen is not present to undergo aerobic respiration, pyruvate will undergo lactic acid
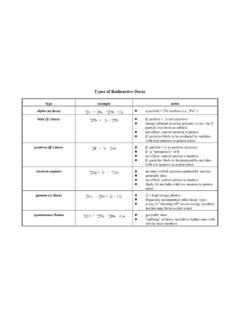
Types of Radioactive Decay - Purdue University
www.chem.purdue.eduTypes of Radioactive Decay type example notes alpha (α) decay 23 9 8 2 U 6 4 2 He + 23 9 4 0 Th + 2 0 0γ! α particle = 4 2 He nucleus (i.e., 4 2 He 2+) beta (β-) decay 23 9 4 0 Th 6! 0 1 e + 23 9 4 1 Pa! β-particle = ! 0 1 e (an electron)! energy released in decay process creates the β-particle (not from an orbital)! net effect: convert ...
Experiment 2: Acid / base titration - Purdue University
www.chem.purdue.eduExperiment 2: Acid / base titration cunknown =±62.0 0.5 mM @ 95% confidence level Nikolai Skrynnikov TA: Boone Prentice Section number: 1 25 Jan 2008 (data courtesy of Ike Fehrenbacher, 2004) 1
THEORY OF XRF - Purdue University
www.chem.purdue.eduXRF, and Chapter 4 describes how this physics is applied to spectrometers and their components. Chapter 5 explains how an XRF analysis is done. It describes the process of sample taking, measuring the sample and calculating the composition from the measurement results.
Functional Groups - Purdue University
www.chem.purdue.eduI. Carboxylic Acids • contain a “carboxylic acid” group – a carbonyl (C=O) group bonded to a hydroxyl group at the carbonyl carbon atom • note that in condensed structural formulas, the carboxylic acid group may be written as –COOH
Purdue University: Department of Chemistry: Home
www.chem.purdue.eduSep 21, 2009 · Draw the structure of the peptide LCYRAIDCG as it would exist at pH 6.5 under oxidizing conditions. Be sure to draw any disulfide bonds that could form. EH — O-NA- Label the N-terminus of the peptide you drew in 3B above with a V. Label the C-terminus of the peptide you drew in 3B above with a Calculate the net charge of LCYRAIDG at pH 1.5.
Related documents
Lecture 3: Enzyme kinetics - School of Informatics ...
www.inf.ed.ac.uk• The pH optimum for enzyme activity is generally close to the pH of the environment/cell location natural for the enzyme • Pepsin (a) is a stomach enzyme that hydrolyzes peptide bonds allowing the digestion of proteins (e.g. in meat and fish). It has a very acidic optimum of pH 1.6, similar to the stomach environment.
Enzymes and Their Functions - Activity Sheets
climb.bme.cornell.edureaction between an enzyme and a substrate is complete, the substrate is changed to a product while the enzyme remains unchanged. The rate of the reaction between an enzyme and a substrate can be affected by different factors. Some of the factors that can affect enzyme activity are temperature, pH,
Enzyme Inhibition: Mechanisms and Scope - IntechOpen
cdn.intechopen.comThe enzyme inhibitors are low molecular weight chemical compounds. They can reduce or completely inhibit the enzyme catalytic activity either reversibly or permanently (irreversibly). Inhibitor can modify one amino acid, or several side chain(s) required in enzyme catalytic activity. To protect enzyme catalytic site from any change, ligand binds
Lab 2. Enzyme Action Effect of Enzyme Concentration ...
instruction.greenriver.eduby the enzyme catalase at various enzyme concentrations, pH, and temperature values. • Define enzyme, catalyst, active site, substrate, activation energy, product, denaturation and explain how enzymes function. • Predict the effects of varying environmental conditions such as pH and temperature on enzyme structure and activity.
Enzyme Assay Units - Deerland Probiotics and Enzymes
www.deerland.comFLU:β-glucosidase activity unit is defined as the One quantity of enzyme that will liberate p-nitrophenol at the rate of 1nmol per minute under the conditions of the assay (pH 7.5 and 30°C). The assay is based on an 8 minute hydrolysis of p-nitrophenyl-β-D-glucopyranoside, in which the liberated p-nitrophenol is measured spectrophotometrically.
LAB Enzymatic Activity of Lactase - Pottsgrove School District
www.pgsd.orgEnzymatic Activity of Lactase OBJECTIVE The purpose of this lab is to explore the properties of the enzyme lactase. By the end of this lab, you should understand the biological function of enzymes and why they are important in the body. You should also examine the specificity of an enzyme to a specific substrate. INTRODUCTION
Regulation of Enzyme Activity
alpha.chem.umb.eduEnzyme Activity Manickam Sugumaran Professor of Biology U.Mass - Boston Boston, MA 02125 The Theme of This Lecture Regulation of Enzyme Activity at Protein Level. 1. Covalent modification. 2. Noncovalent (allosteric) regulation 3. Protein degradation (will not be considered). Transcriptional regulation Translational regulation
Activity, Regulations, Enzymes, Enzyme activity, Regulation of enzyme activity